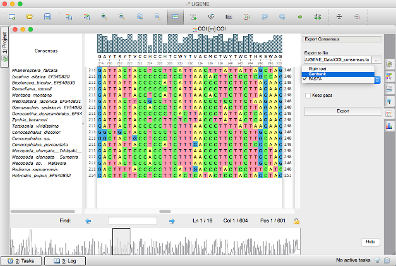
In contrast to Clustal Omega, which replaces amino acids by numbers to speed up pairwise similarity score calculations, ClustalW builds the guide tree based on real pairwise alignments. Now we will make the alignment using ClustalW2.
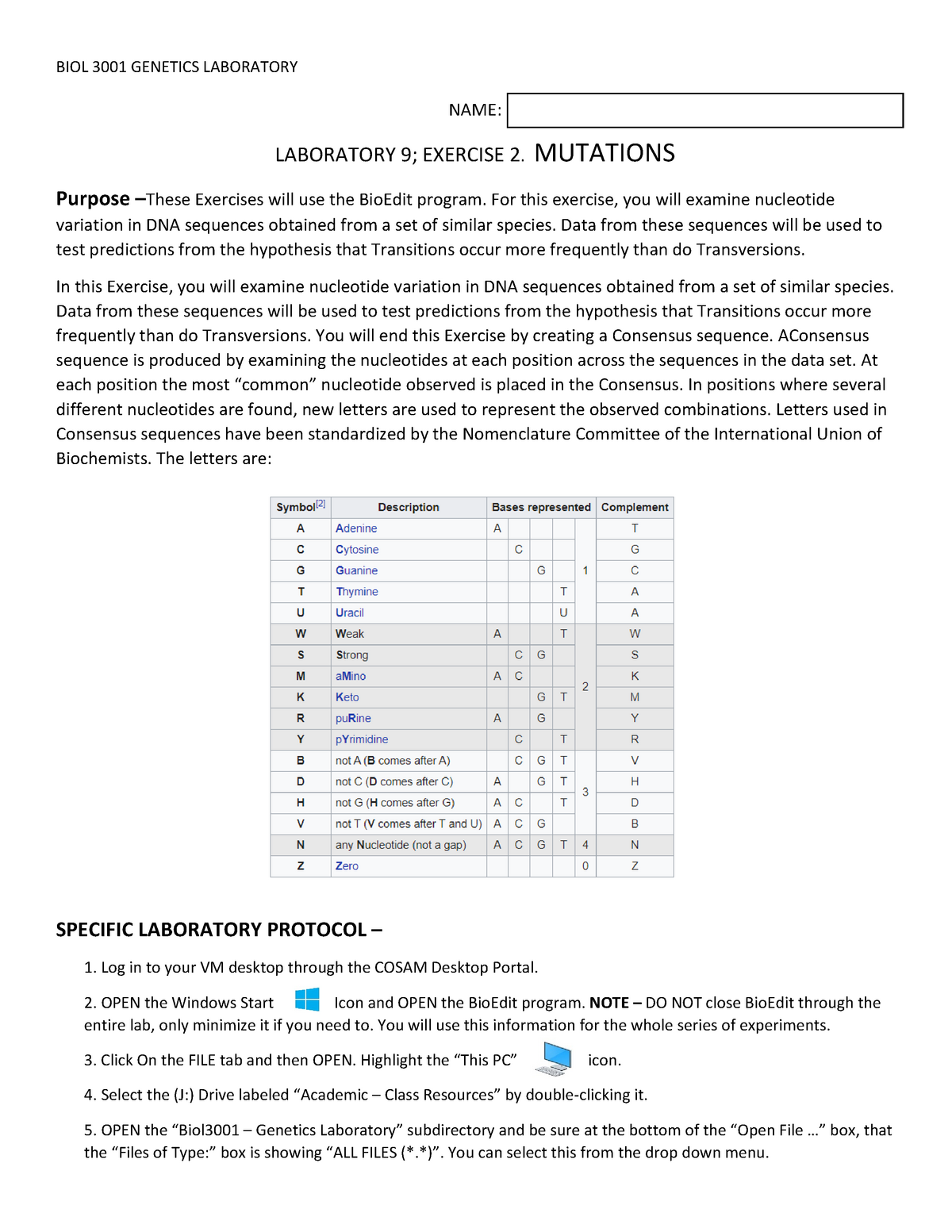
#GETTING CONSENSUS IN BIOEDIT DOWNLOAD#
You can download the sequences in fasta format.

We are first going to use different tools for the toy example below:
#GETTING CONSENSUS IN BIOEDIT MANUAL#
Some MSA tools allow manual editing of alignments, which will be discussed in the next exercise. Let's discover how some of these programs behave, and which suit your needs best! As always, EBI provides a nice collection of web-based alignment tools but you can also make MSAs in Ugene. Fortunately, you can also find tools to convert MSA formats. Several multiple sequence algorithms exist, each with their own program and format.